Normal/gaussian/bell curve distributions and more to illustrate probability density functions (pdfs), sigma bands (68-95-99 rule), test statistics, critical regions (accept/reject), type-I and type-II errors (alpha, beta), null hypothesis tests, p-value, low sensitivity, bias & systematic error, S+B vs. B-only hypotheses, upper limits and the CLs method.
Inspired by Glen Cowen’s CERN lectures and this Higgs physics course by Mauro Donega at the UZH & ETHZ. Presented in this talk on the statistical method in particle physics.
Also see critical regions and these sets of critical regions.
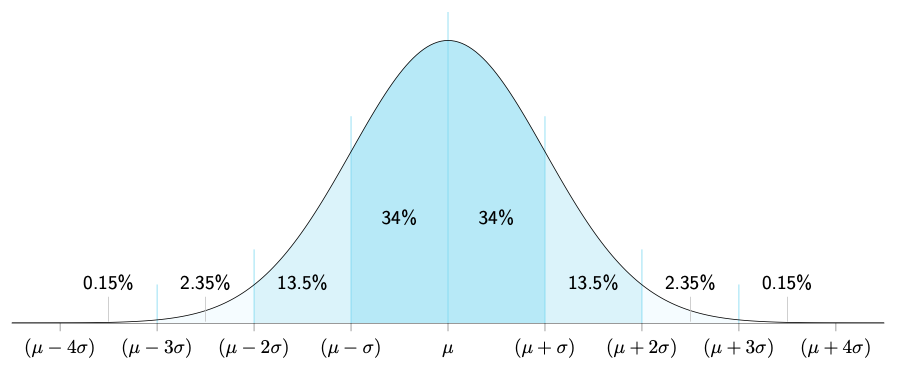
Probability from gaussian probability density functions (pdfs):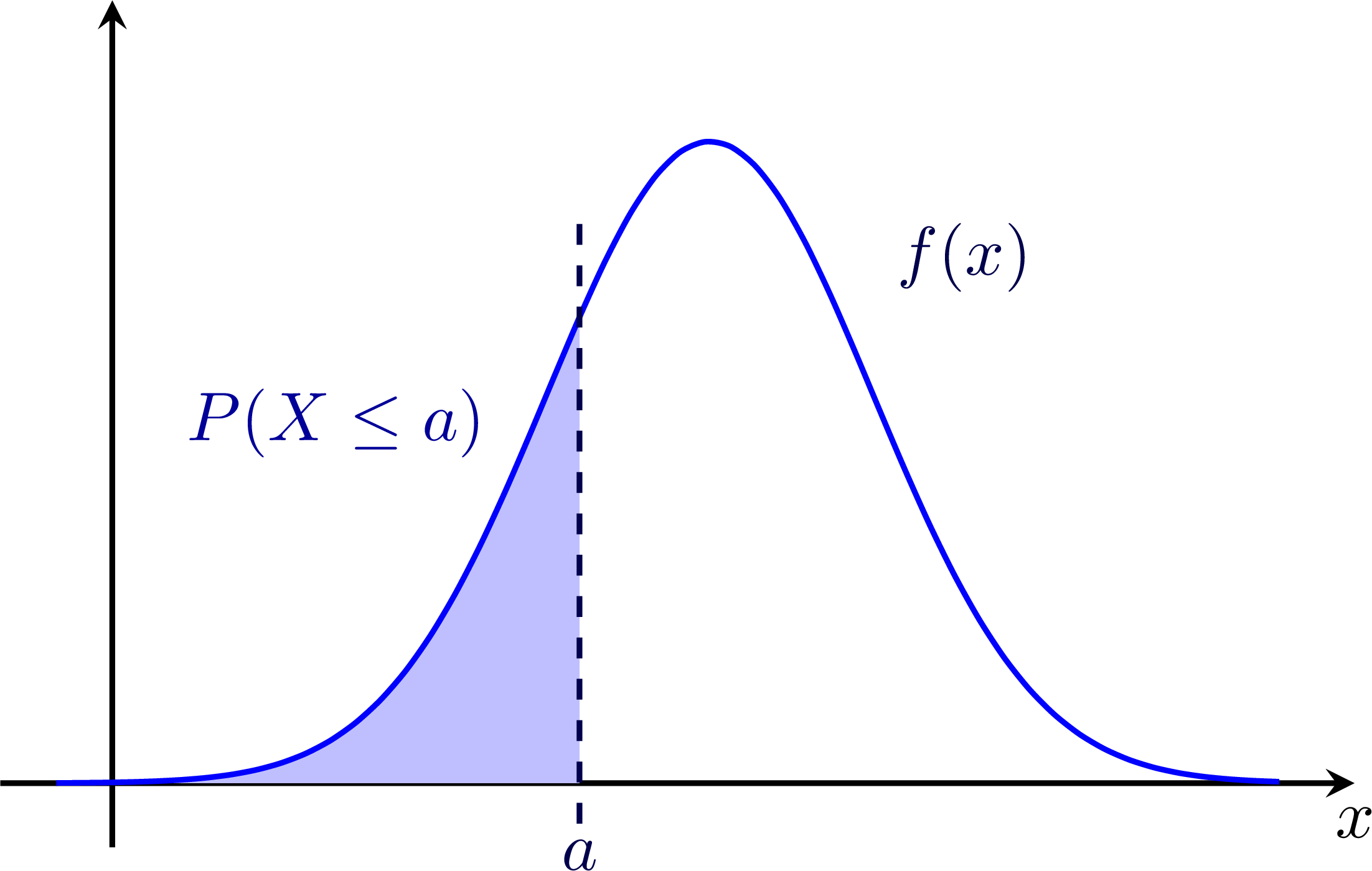 The 68-95-99 rule:
The 68-95-99 rule: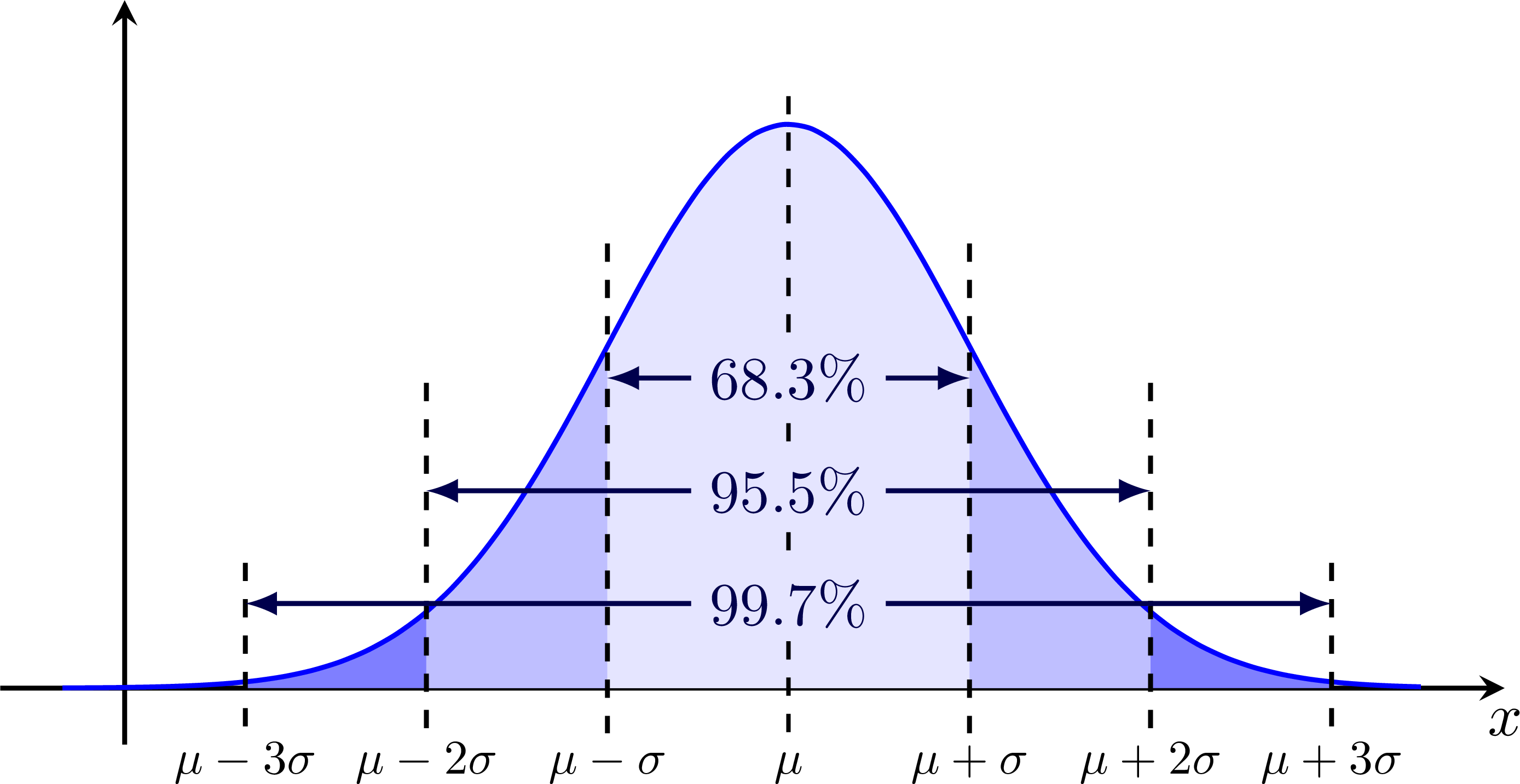 Sigma (standard deviation) bands for 'Brazilian flag' plots:
Sigma (standard deviation) bands for 'Brazilian flag' plots: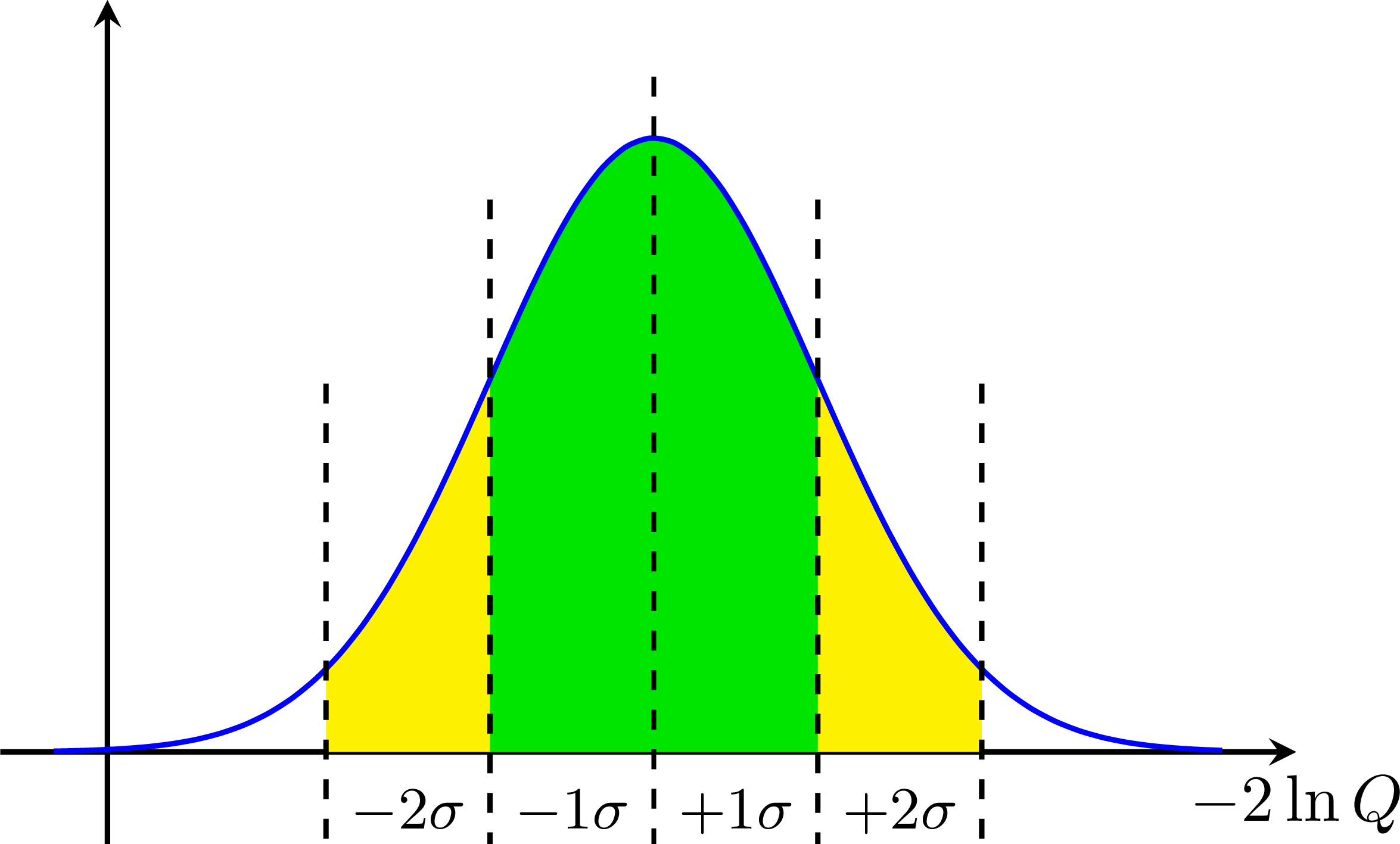 Hypothesis testing with a test statistics t to define a critical region (accept/reject) with type-I (alpha) and type-II errors (beta):
Hypothesis testing with a test statistics t to define a critical region (accept/reject) with type-I (alpha) and type-II errors (beta):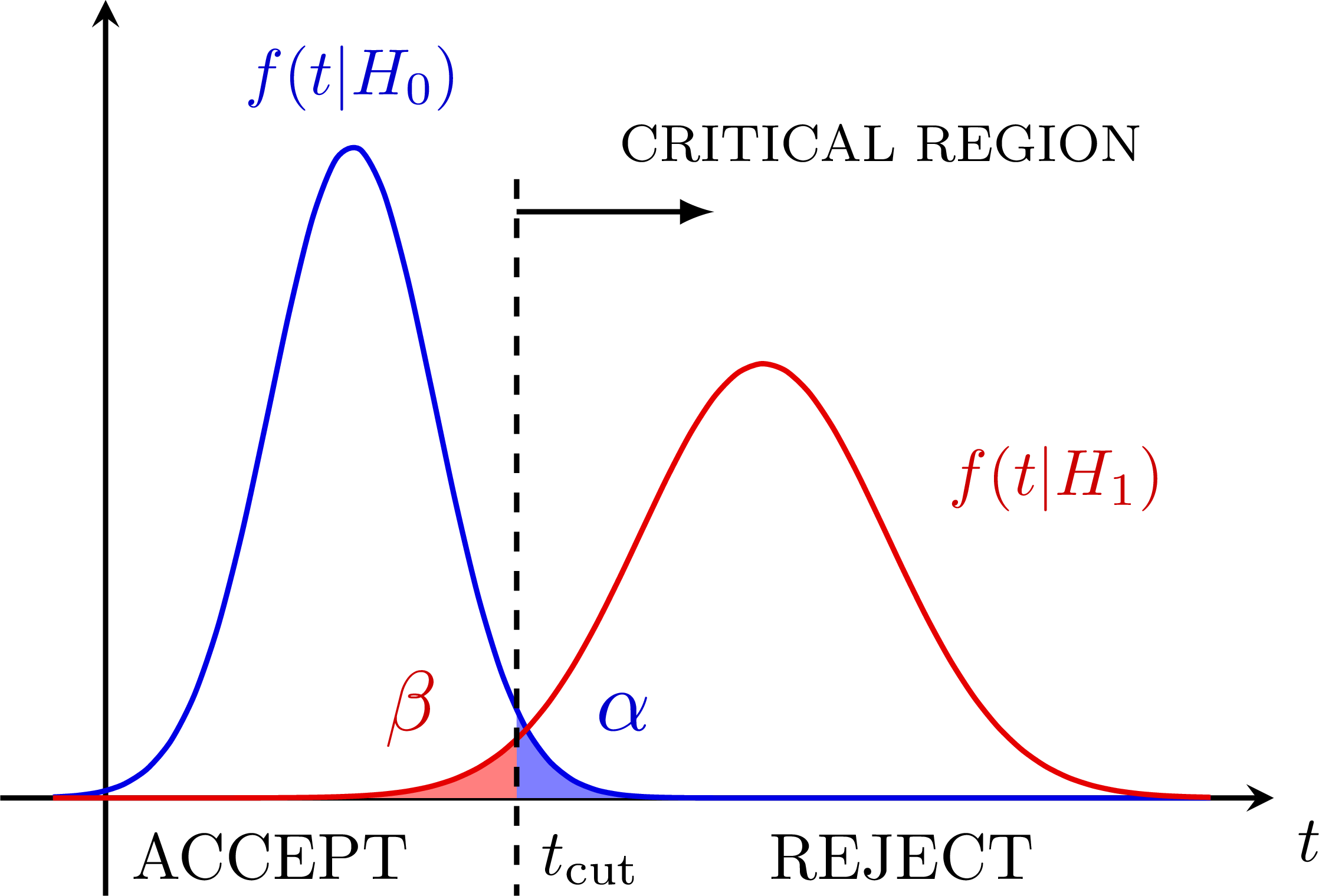 P-value (for a null-hypothesis):
P-value (for a null-hypothesis):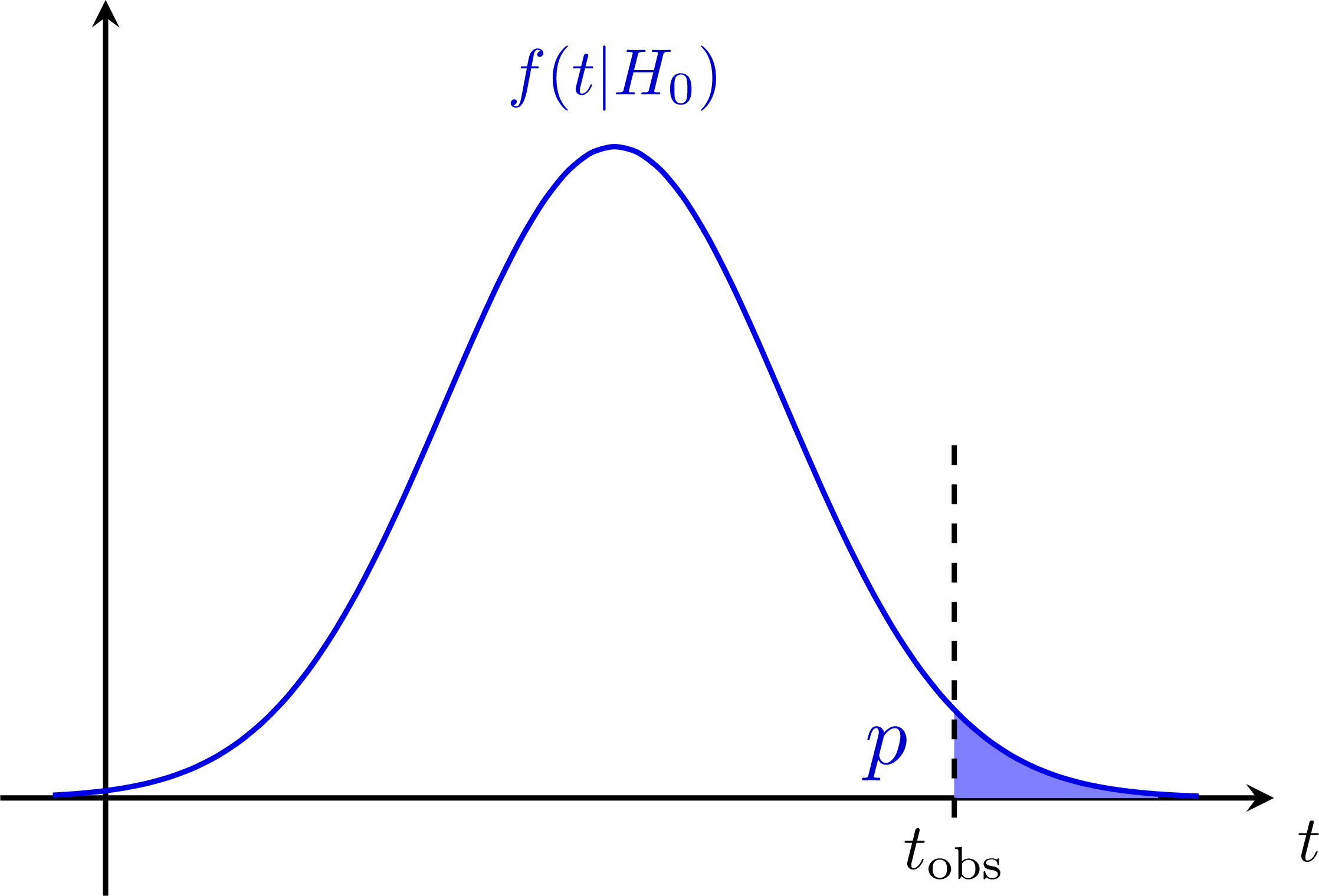 P-value for a B-only model (signal strength μ = 0):
P-value for a B-only model (signal strength μ = 0):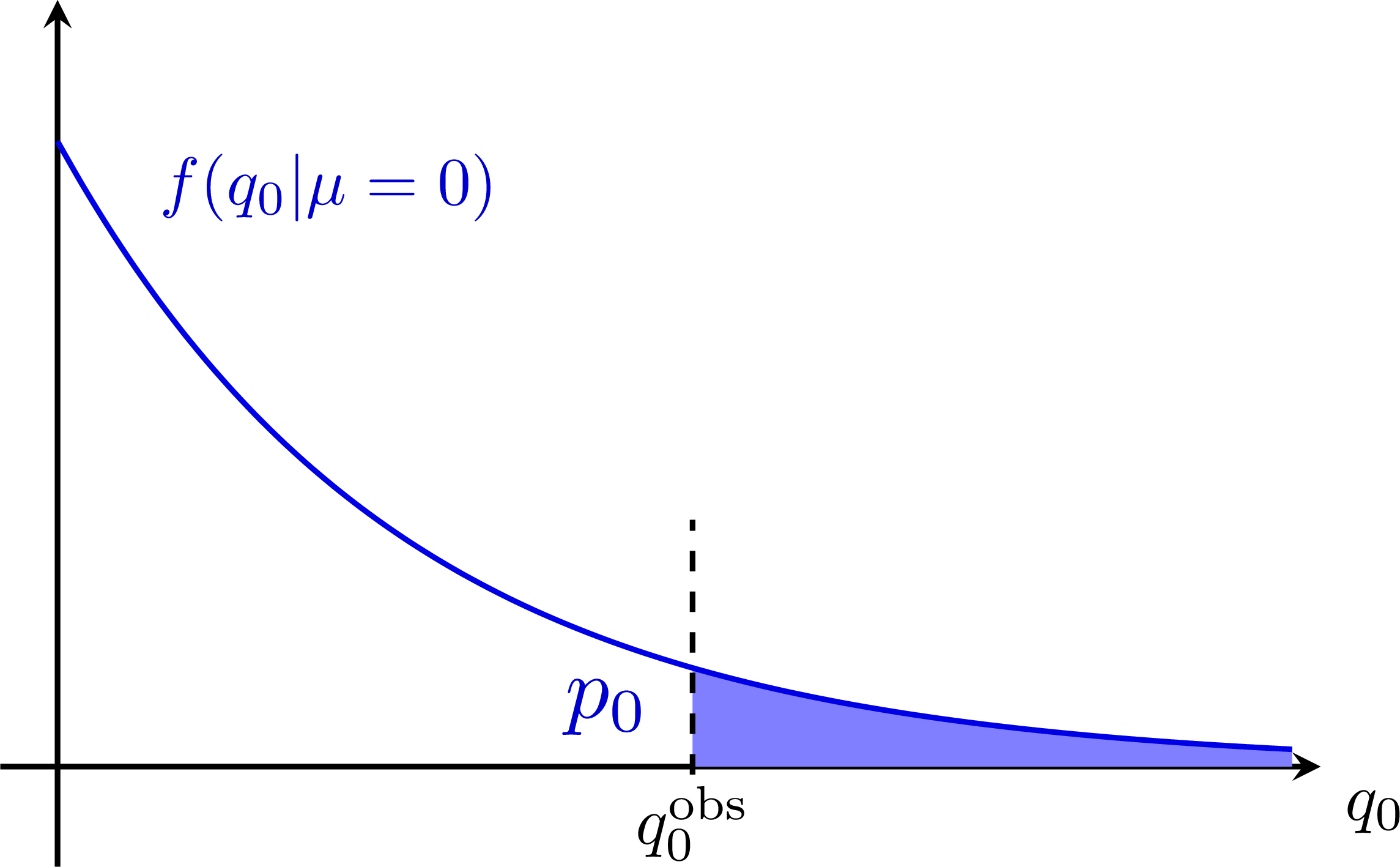
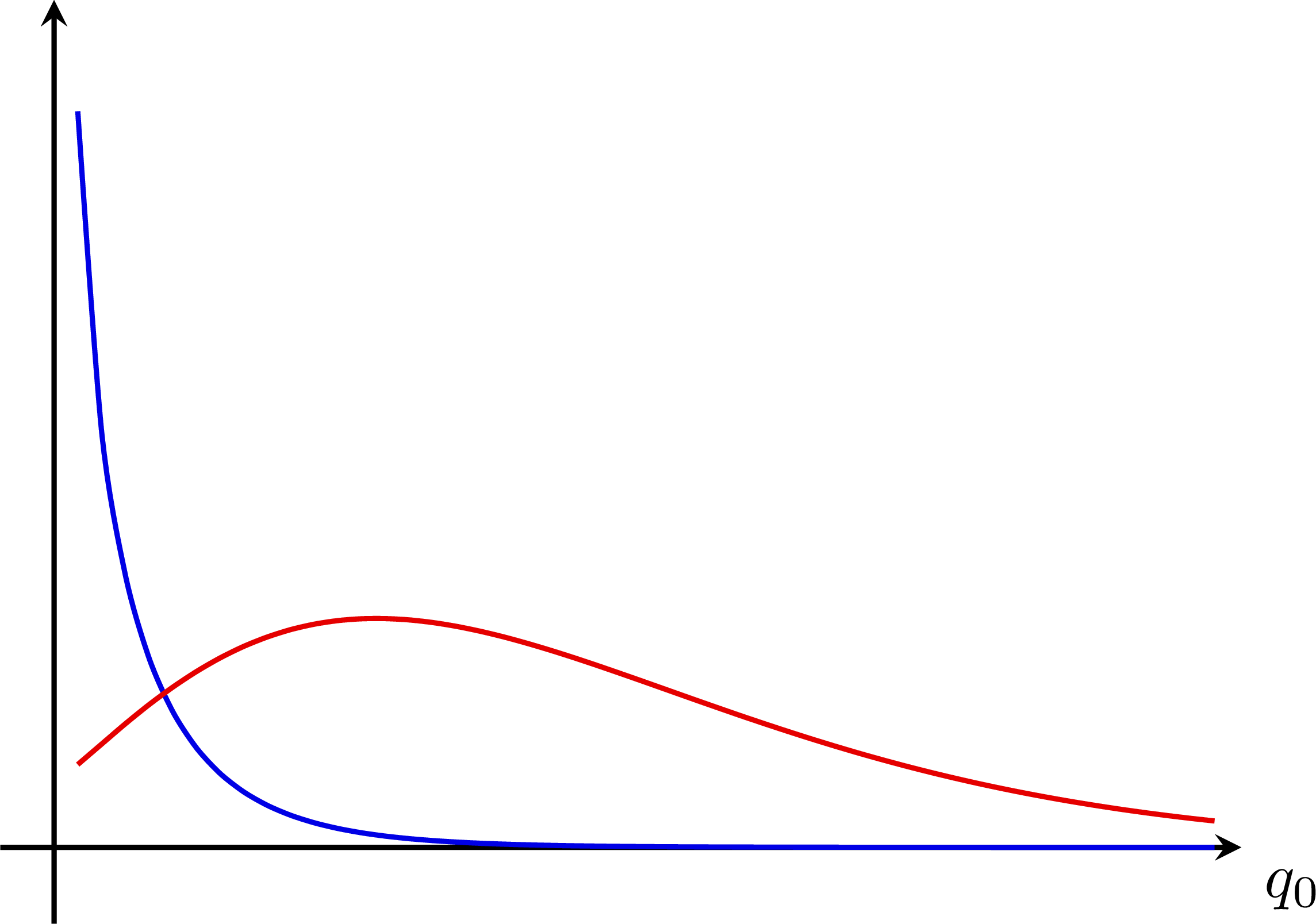 P-value for an S+B and B-only hypothesis:
P-value for an S+B and B-only hypothesis: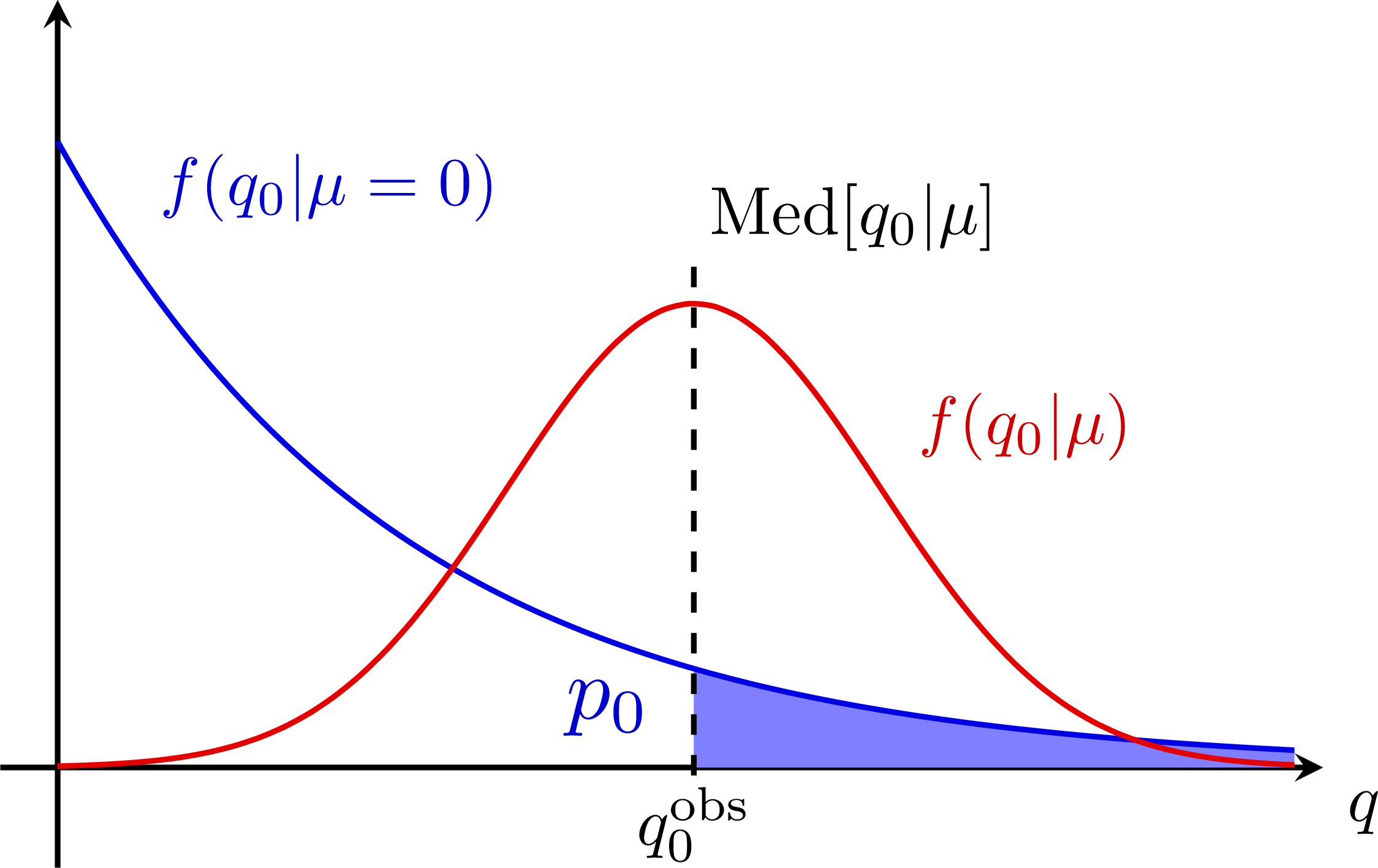 Bias & systematic error from nuisance parameters:
Bias & systematic error from nuisance parameters: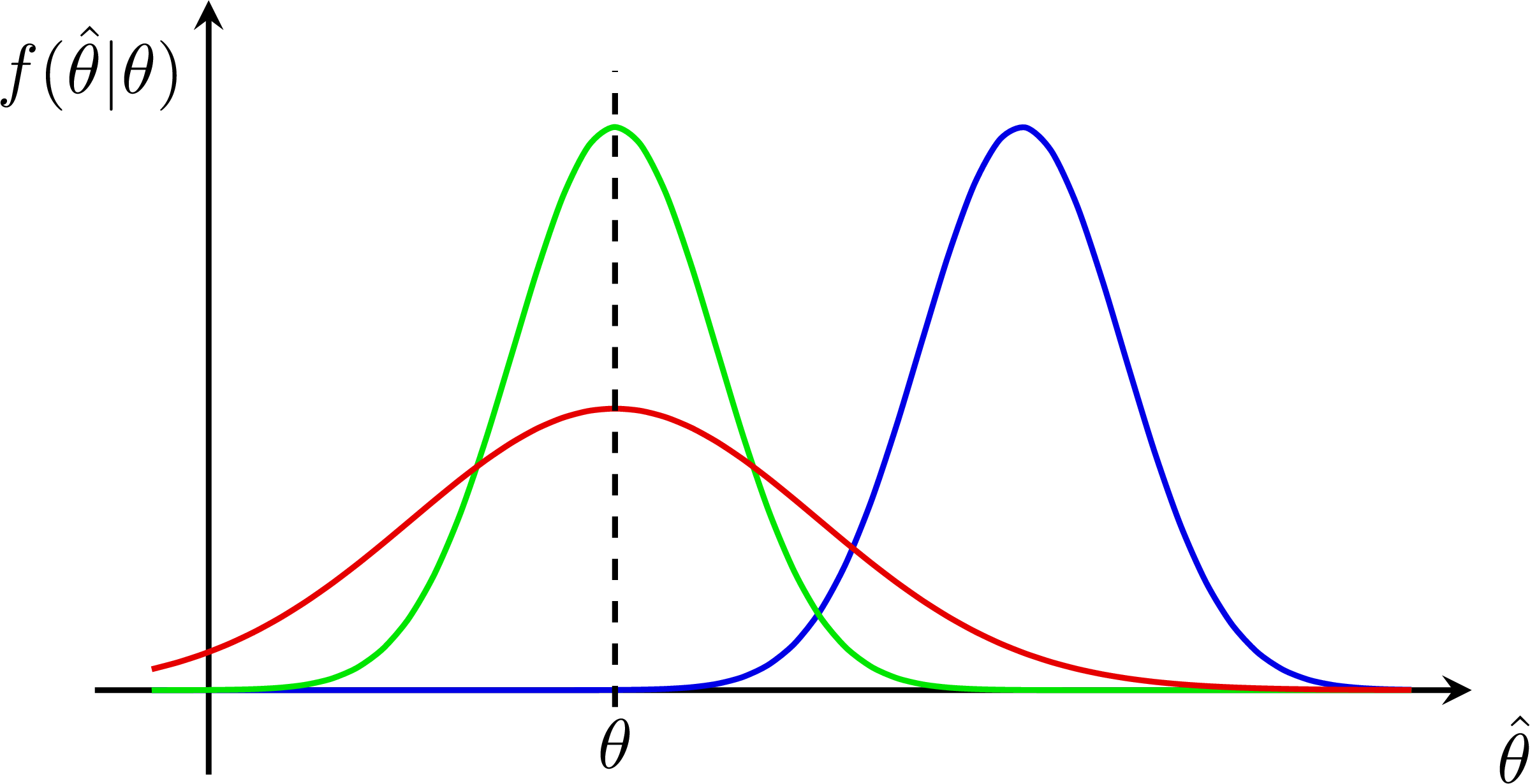 Z-scores for a normal distribution:
Z-scores for a normal distribution: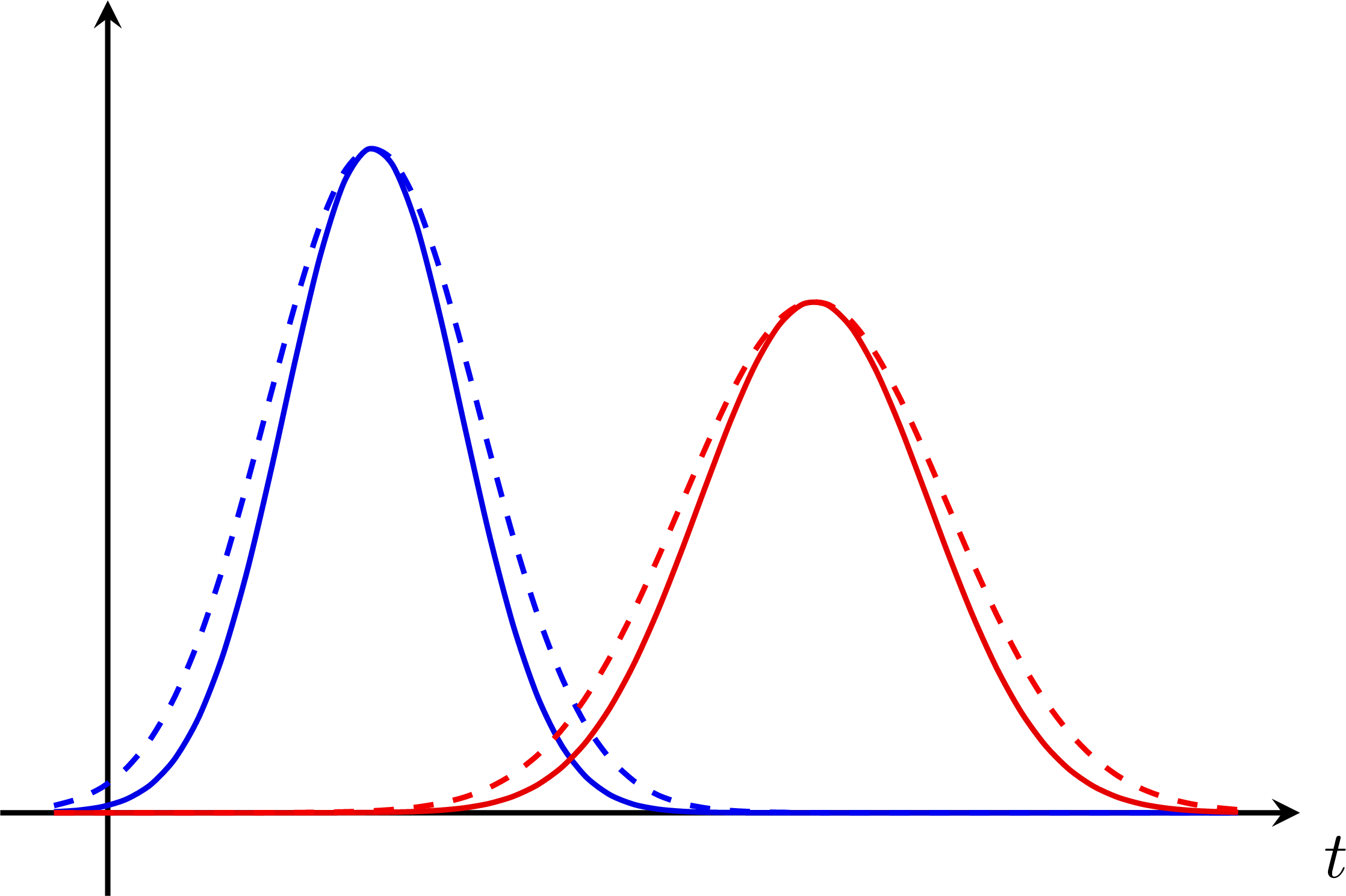 Hypothesis testing with null-hypothesis H0:
Hypothesis testing with null-hypothesis H0: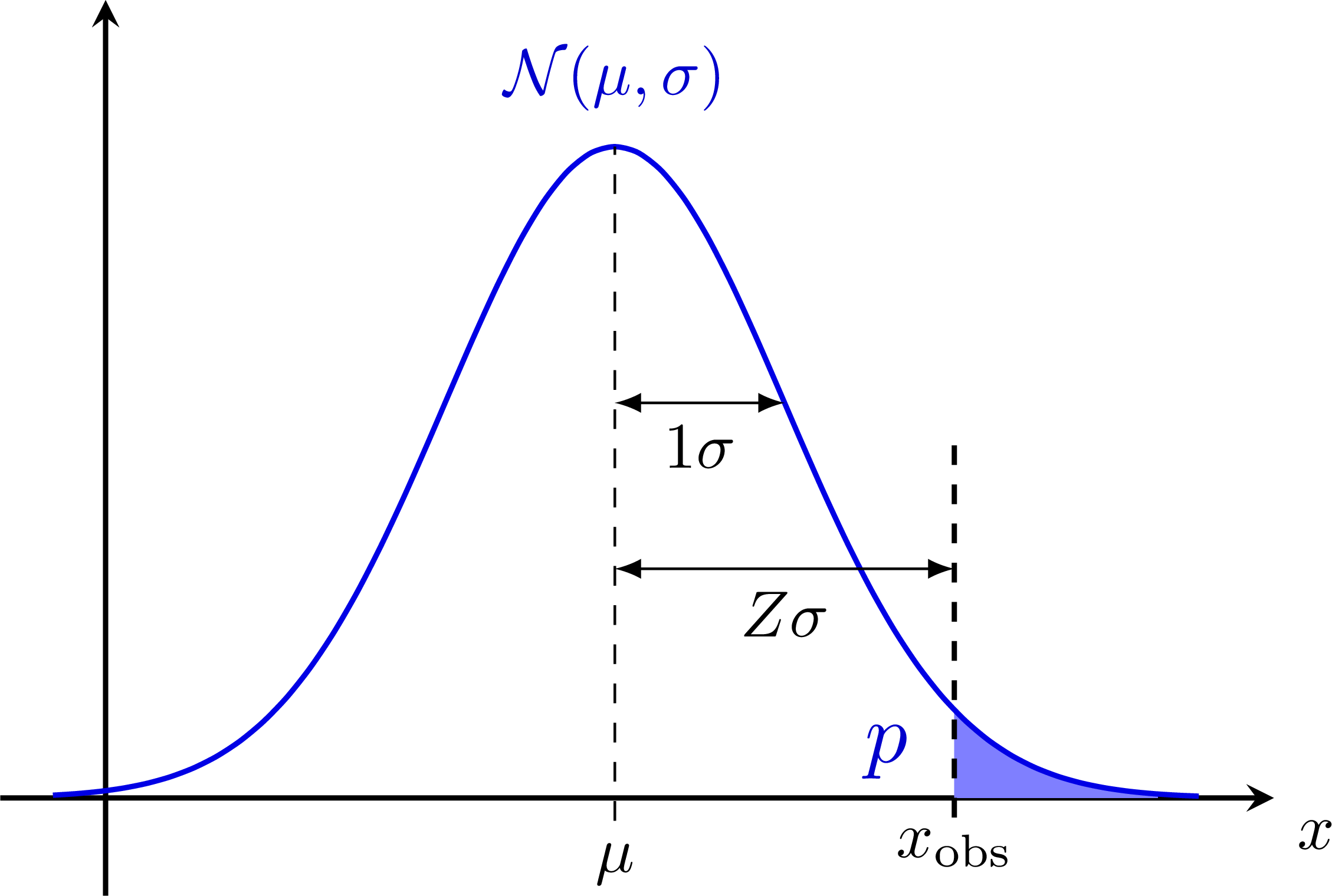 Hypothesis testing with two gaussians:
Hypothesis testing with two gaussians: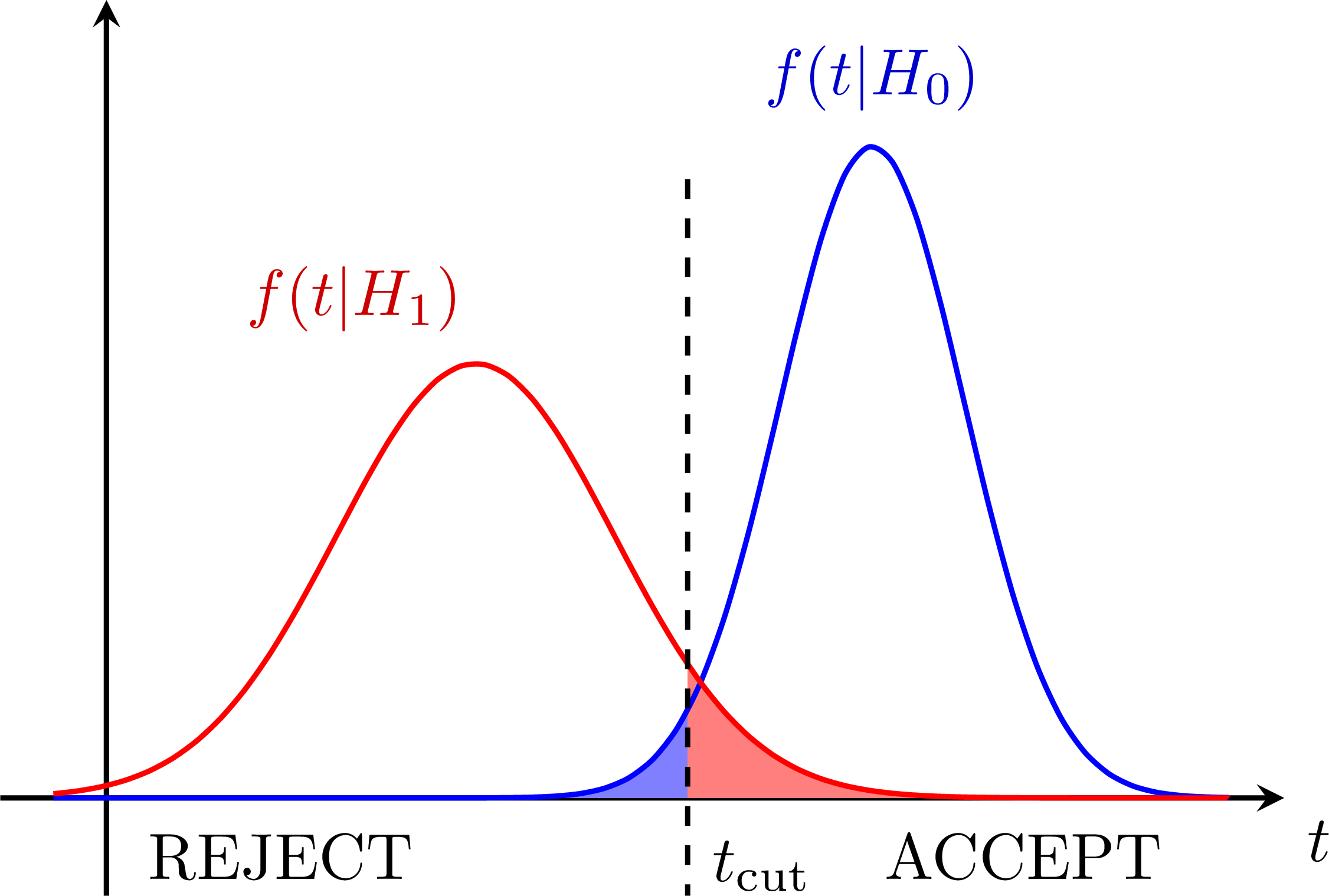 Test power:
Test power: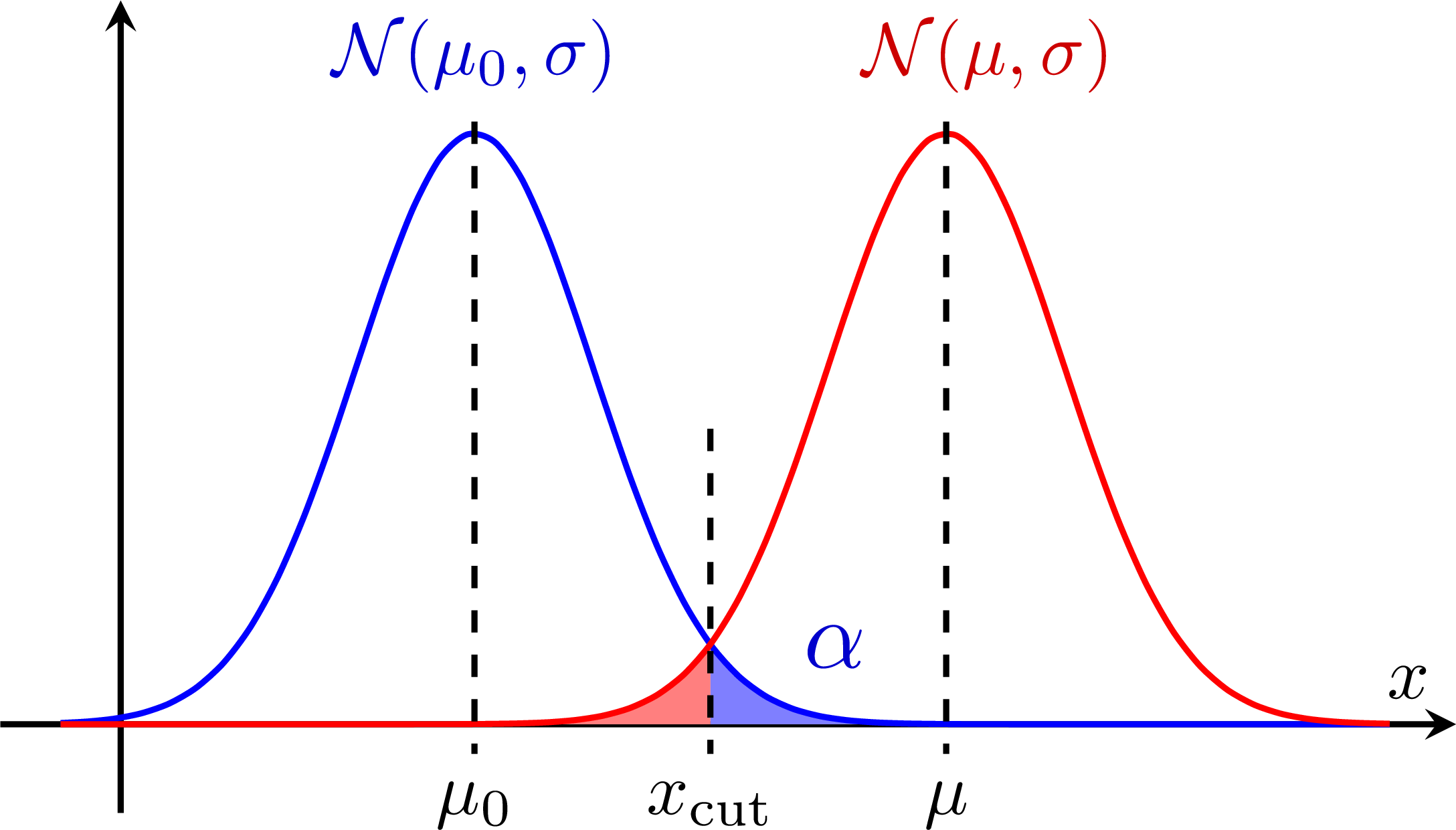 Low power/sensitivity:
Low power/sensitivity: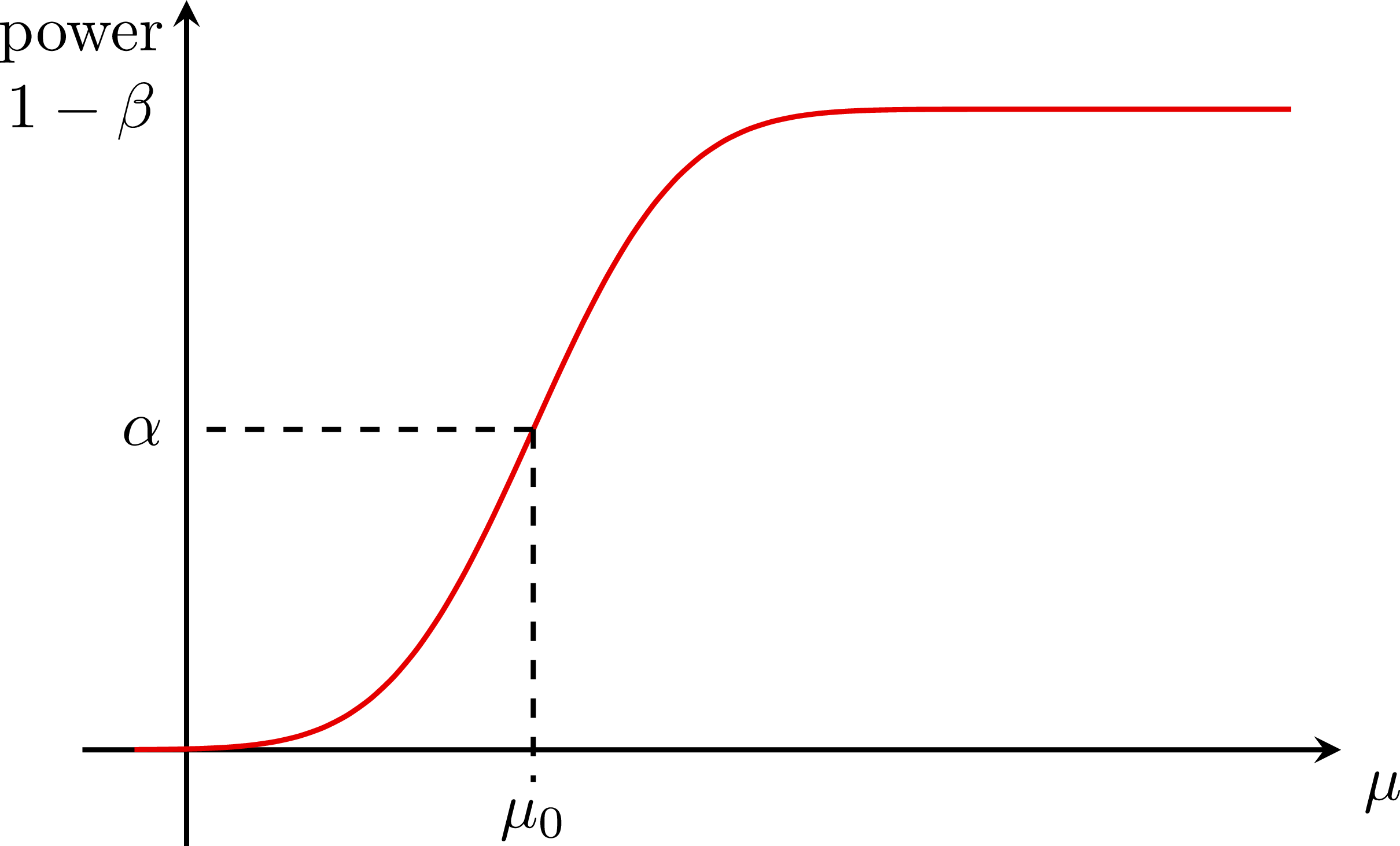 Better power:
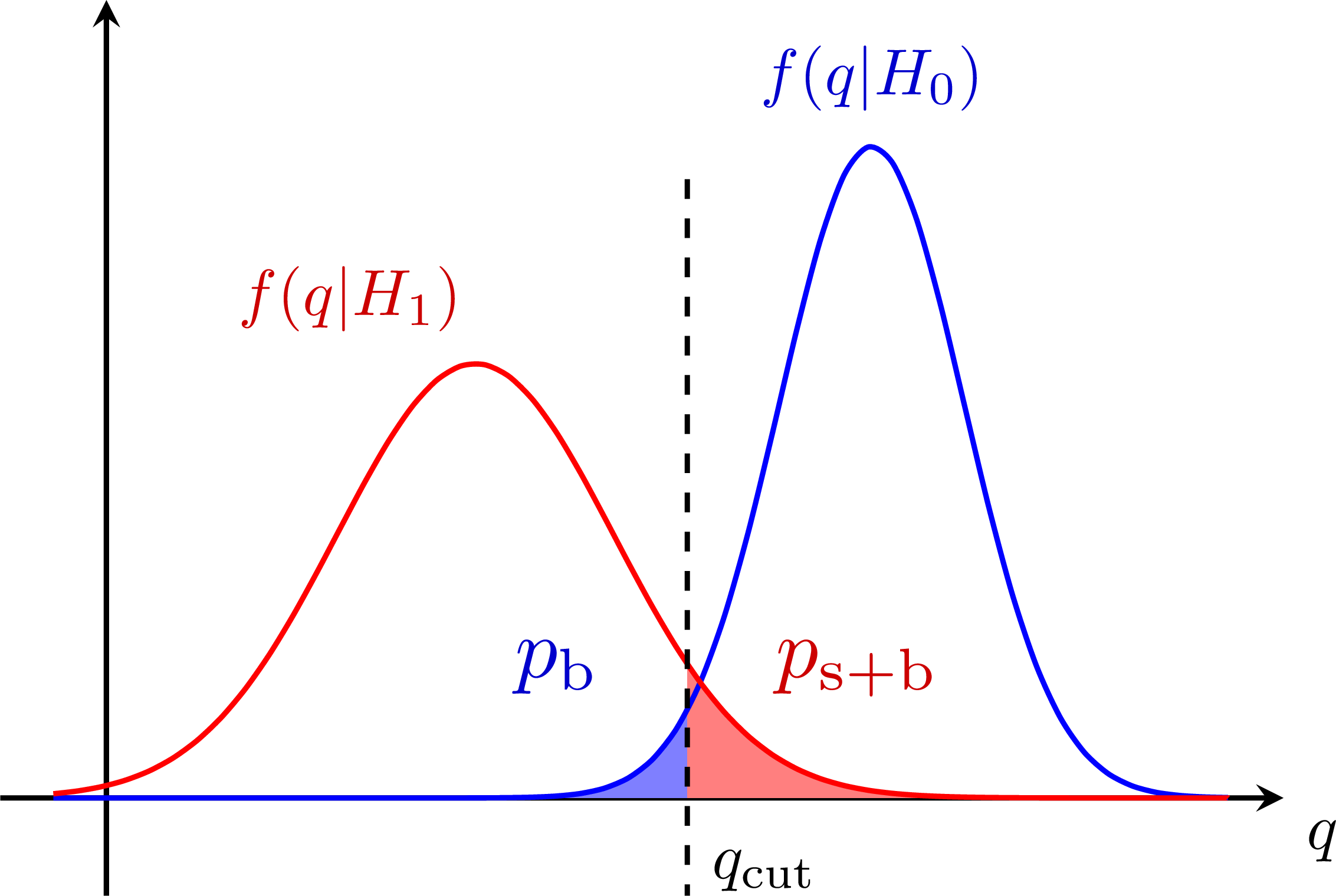
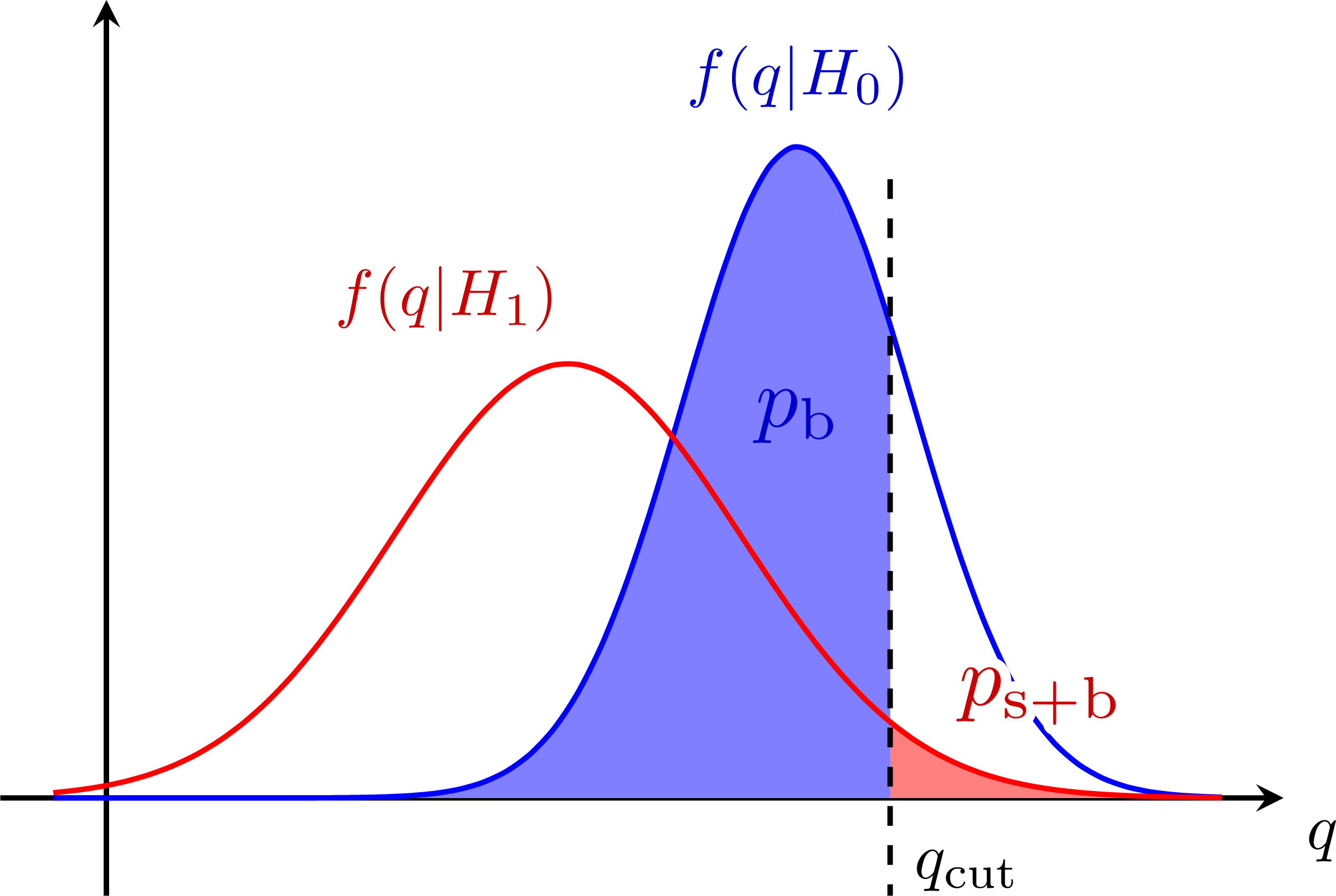
Better power: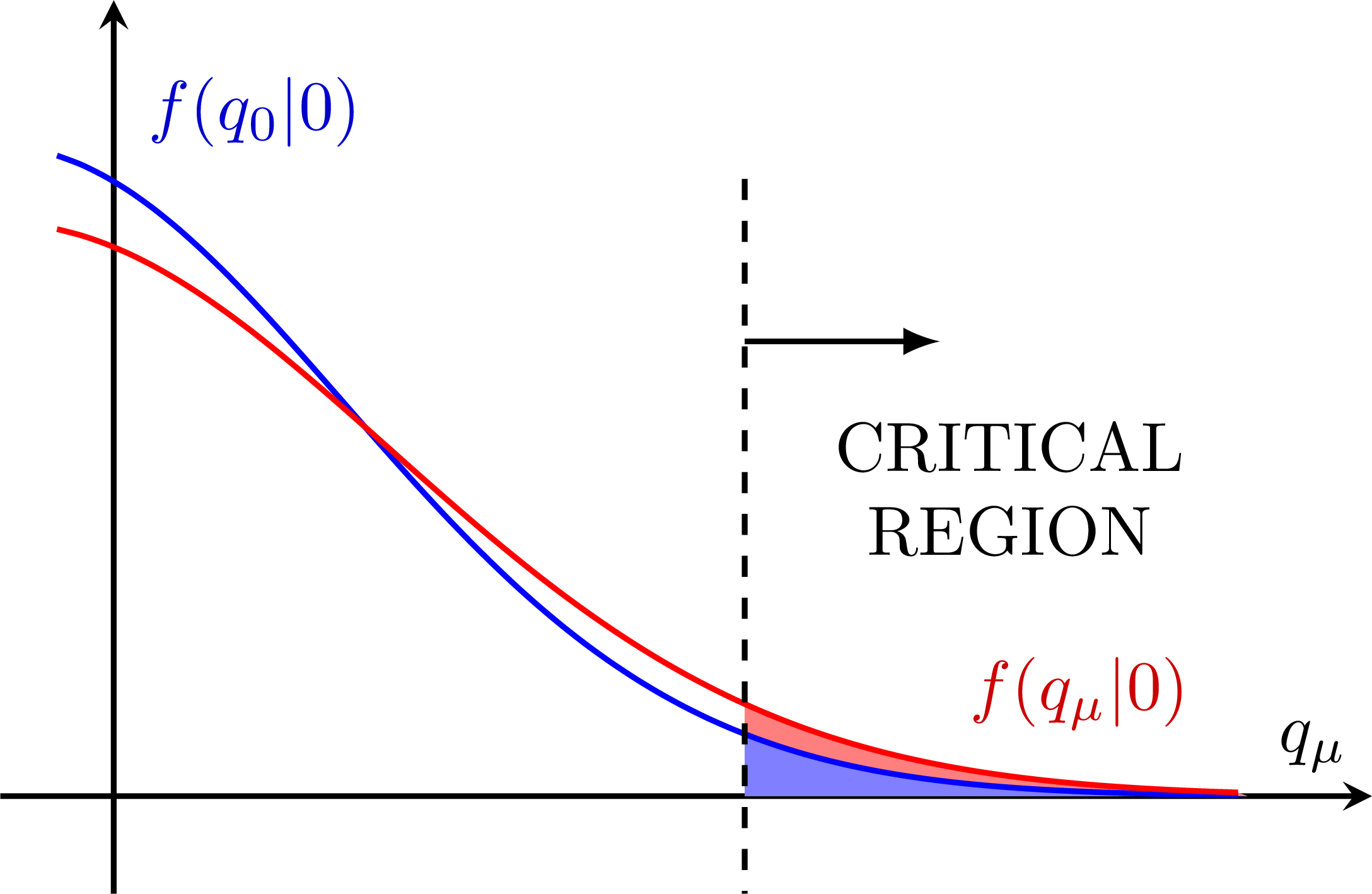 S+B and B-only p-values for the CLs method:
S+B and B-only p-values for the CLs method: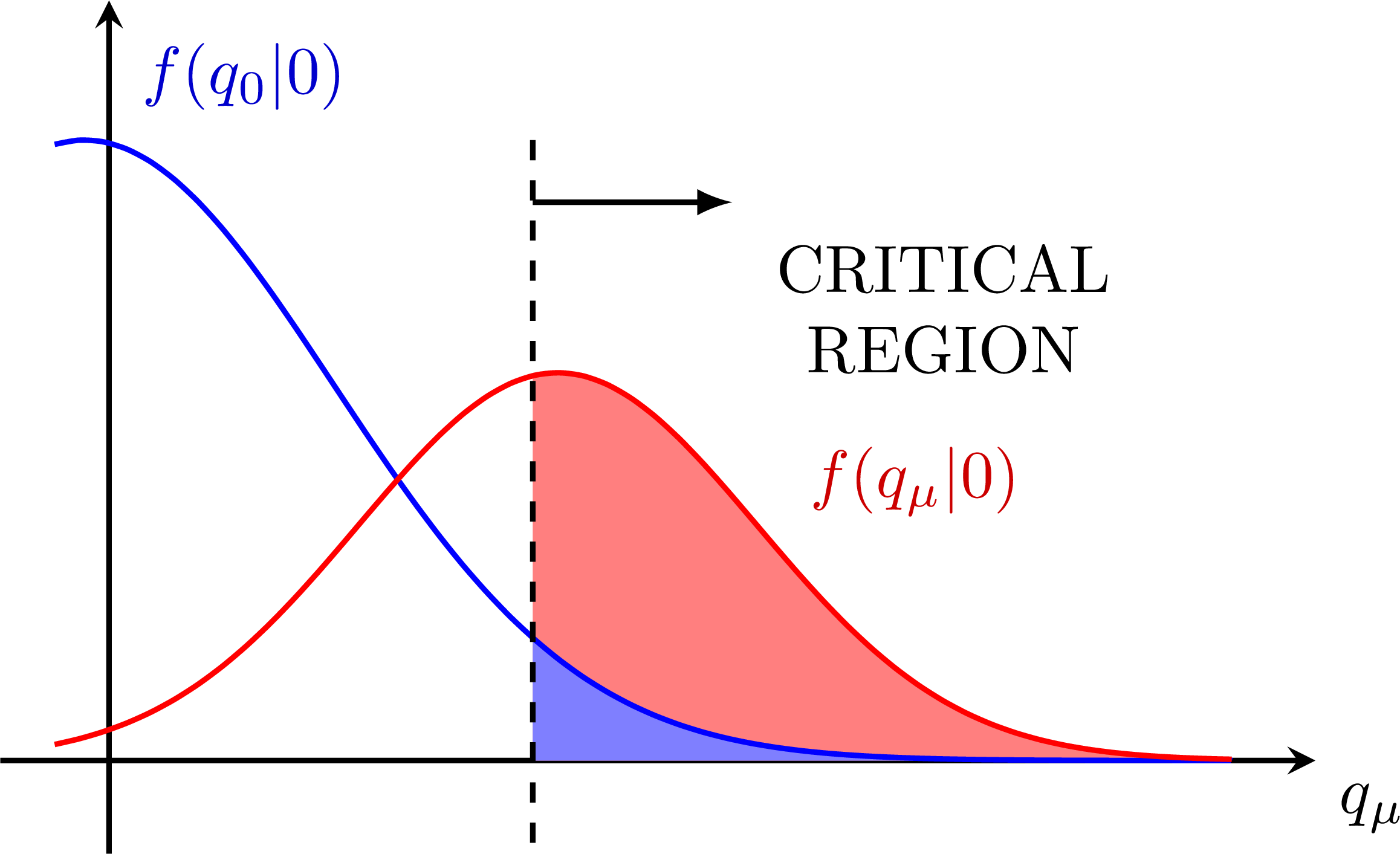
Edit and compile if you like:
% Author: Izaak Neutelings (August, 2017)
\documentclass[border=3pt,tikz]{standalone} %[dvipsnames]
\usepackage{amsmath} % for \dfrac
\usepackage{tikz}
\tikzset{>=latex} % for LaTeX arrow head
\usepackage{pgfplots} % for the axis environment
\usepackage{xcolor}
\usepackage[outline]{contour} % halo around text
\contourlength{1.2pt}
\usetikzlibrary{positioning,calc}
\usetikzlibrary{backgrounds}% required for 'inner frame sep'
%\usepackage{adjustbox} % add whitespace (trim)
% define gaussian pdf and cdf
\pgfmathdeclarefunction{gauss}{3}{%
\pgfmathparse{1/(#3*sqrt(2*pi))*exp(-((#1-#2)^2)/(2*#3^2))}%
}
\pgfmathdeclarefunction{cdf}{3}{%
\pgfmathparse{1/(1+exp(-0.07056*((#1-#2)/#3)^3 - 1.5976*(#1-#2)/#3))}%
}
\pgfmathdeclarefunction{fq}{3}{%
\pgfmathparse{1/(sqrt(2*pi*#1))*exp(-(sqrt(#1)-#2/#3)^2/2)}%
}
\pgfmathdeclarefunction{fq0}{1}{%
\pgfmathparse{1/(sqrt(2*pi*#1))*exp(-#1/2))}%
}
\colorlet{mydarkblue}{blue!30!black}
% to fill an area under function
\usepgfplotslibrary{fillbetween}
\usetikzlibrary{patterns}
\pgfplotsset{compat=1.12} % TikZ coordinates <-> axes coordinates
% https://tex.stackexchange.com/questions/240642/add-vertical-line-of-equation-x-2-and-shade-a-region-in-graph-by-pgfplots
% plot aspect ratio
%\def\axisdefaultwidth{8cm}
%\def\axisdefaultheight{6cm}
% number of sample points
\def\N{50}
\begin{document}
% GAUSSIANs: basic properties
\begin{tikzpicture}
\message{Cumulative probability^^J}
\def\B{11};
\def\Bs{3.0};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.1*gauss(\B,\B,\Bs)}};
\def\h{0.07*gauss(\B,\B,\Bs)};
\def\a{\B-0.8*\Bs};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$x$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north},
width=0.7*\textwidth, height=0.55*\textwidth,
y=700pt,
clip=false
]
% PLOTS
\addplot[blue,thick,name path=B] {gauss(x,\B,\Bs)};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0);
\addplot[blue!25] fill between[of=xaxis and B, soft clip={domain=-1:{\a}}];
% LINES
\addplot[mydarkblue,dashed,thick]
coordinates {({\a},{1.2*gauss(\a,\B,\Bs)}) ({\a},{-\h})}
node[mydarkblue,below=-2pt] {$a$};
\node[mydarkblue,above right] at ({\B+\Bs},{1.2*gauss(\B+\Bs,\B,\Bs)}) {$f(x)$};
\node[blue!60!black,above left] at ({0.85*(\a)},{1.0*gauss(0.85*(\a),\B,\Bs)}) {$P(X\leq a)$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: 68-95-99 rule
\begin{tikzpicture}
\message{68-95-99 rule^^J}
\def\B{11};
\def\Bs{3.0};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.1*gauss(\B,\B,\Bs)}};
\def\h{0.08*gauss(\B,\B,\Bs)};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$x$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north},
width=0.85*\textwidth, height=0.55*\textwidth,
y=700pt,
clip=false
]
% PLOTS
\addplot[blue,thick,name path=B] {gauss(x,\B,\Bs)};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[blue!50] fill between[of=xaxis and B, soft clip={domain={\B-3*\Bs}:{\B+3*\Bs}}];
\addplot[blue!25] fill between[of=xaxis and B, soft clip={domain={\B-2*\Bs}:{\B+2*\Bs}}];
\addplot[blue!10] fill between[of=xaxis and B, soft clip={domain={\B-1*\Bs}:{\B+1*\Bs}}];
% LINES
\addplot[black,dashed,thick]
coordinates {({\B-3*\Bs},{20*gauss(\B-3*\Bs,\B,\Bs)}) ({\B-3*\Bs},{-\h})}
node[below=-3pt,scale=0.8] {\strut$\mu-3\sigma$};
\addplot[black,dashed,thick]
coordinates {({\B-2*\Bs},{4*gauss(\B-2*\Bs,\B,\Bs)}) ({\B-2*\Bs},{-\h})}
node[below=-3pt,scale=0.8] {\strut$\mu-2\sigma$};
\addplot[black,dashed,thick]
coordinates {({\B-1*\Bs},{1.3*gauss(\B-\Bs,\B,\Bs)}) ({\B-1*\Bs},{-\h})}
node[below=-3pt,scale=0.8] at ({\B-\Bs},{-\h}) {\strut$\mu-\sigma$};
\addplot[black,dashed,line width=0.7pt]
coordinates {(\B,{1.05*gauss(\B,\B,\Bs)}) (\B,{-\h})}
node[below=-3pt,scale=0.8] {\strut$\mu$};
\addplot[black,dashed,thick]
coordinates {({\B+1*\Bs},{1.3*gauss(\B+\Bs,\B,\Bs)}) ({\B+1*\Bs},{-\h})}
node[below=-3pt,scale=0.8] at ({\B+\Bs},{-\h}) {\strut$\mu+\sigma$};
\addplot[black,dashed,thick]
coordinates {({\B+2*\Bs},{4*gauss(\B+2*\Bs,\B,\Bs)}) ({\B+2*\Bs},{-\h})}
node[below=-3pt,scale=0.8] at ({\B+2*\Bs},{-\h}) {\strut$\mu+2\sigma$};
\addplot[black,dashed,thick]
coordinates {({\B+3*\Bs},{20*gauss(\B+3*\Bs,\B,\Bs)}) ({\B+3*\Bs},{-\h})}
node[below=-3pt,scale=0.8] at ({\B+3*\Bs},{-\h}) {\strut$\mu+3\sigma$};
% AREAS
\addplot[<->,mydarkblue,thick]
coordinates {({\B-\Bs},{.55*gauss(\B,\B,\Bs)}) ({\B+\Bs},{.55*gauss(\B,\B,\Bs)})};
\addplot[<->,mydarkblue,thick]
coordinates {({\B-2*\Bs},{.35*gauss(\B,\B,\Bs)}) ({\B+2*\Bs},{.35*gauss(\B,\B,\Bs)})};
\addplot[<->,mydarkblue,thick]
coordinates {({\B-3*\Bs},{.15*gauss(\B,\B,\Bs)}) ({\B+3*\Bs},{.15*gauss(\B,\B,\Bs)})};
\node[mydarkblue,fill=blue!10,inner xsep=3,inner ysep=1,scale=1]
at (\B,{.55*gauss(\B,\B,\Bs)}) {68.3\%};
\node[mydarkblue,fill=blue!10,inner xsep=3,inner ysep=2,scale=1]
at (\B,{.35*gauss(\B,\B,\Bs)}) {95.5\%};
\node[mydarkblue,fill=blue!10,inner xsep=3,inner ysep=2,scale=1]
at (\B,{.15*gauss(\B,\B,\Bs)}) {99.7\%};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: error bands
\begin{tikzpicture}
\message{Error bands^^J}
\def\B{10};
\def\Bs{3.0};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$-2\ln Q$, %q
every axis x label/.style={at={(current axis.right of origin)},anchor=north},
width=0.7*\textwidth, height=0.5*\textwidth,
y=700pt
]
% PLOTS
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[black,dashed,thick]
coordinates {({\B-2*\Bs}, {0.60*gauss(\B,\B,\Bs)}) ({\B-2*\Bs}, \ymin)};
\addplot[black,dashed,thick]
coordinates {({\B-1*\Bs}, {0.90*gauss(\B,\B,\Bs)}) ({\B-1*\Bs}, \ymin)};
\addplot[black,dashed,line width=0.7pt]
coordinates {(\B, {1.10*gauss(\B,\B,\Bs)}) (\B,\ymin)};
\addplot[black,dashed,thick]
coordinates {({\B+1*\Bs}, {0.90*gauss(\B,\B,\Bs)}) ({\B+1*\Bs}, \ymin)};
\addplot[black,dashed,thick]
coordinates {({\B+2*\Bs}, {0.60*gauss(\B,\B,\Bs)}) ({\B+2*\Bs}, \ymin)};
\node[above=-2pt] at ({\B-1.5*\Bs},\ymin) {\small$-2\sigma$};
\node[above=-2pt] at ({\B-0.5*\Bs},\ymin) {\small$-1\sigma$};
\node[above=-2pt] at ({\B+0.5*\Bs},\ymin) {\small$+1\sigma$};
\node[above=-2pt] at ({\B+1.5*\Bs},\ymin) {\small$+2\sigma$};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[black!0!yellow] fill between[of=xaxis and B, soft clip={domain={\B-2*\Bs}:{\B+2*\Bs}}];
\addplot[black!10!green] fill between[of=xaxis and B, soft clip={domain={\B-1*\Bs}:{\B+1*\Bs}}];
% LABELS
%\node[above=0pt, black!20!blue] at (\B,{1.05*gauss(\B,\B,\Bs)}) {$f(x|\text{b})$};
%\node[] at ($(\ymin,0)!0.5!(\ymin,0)$) {$+2\sigma$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: confidence level
\begin{tikzpicture}
\message{Confidence level^^J}
\def\q{5};
\def\B{3};
\def\S{8};
\def\Bs{1.0};
\def\Ss{1.5};
\def\xmax{\S+3.2*\Ss};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$t$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=250pt
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {gauss(x,\B,\Bs)};
\addplot[name path=S,thick,black!10!red ] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q, {0.95*gauss(\B,\B,\Bs)}) (\q, \ymin)}
node[below=2pt,anchor=south west] {$t_\text{cut}$};
\draw[->,thick]
(\q,{0.90*gauss(\B,\B,\Bs)}) -- ({\q+1.6*\Ss},{0.90*gauss(\B,\B,\Bs)})
node[above=4pt] {\footnotesize\qquad\qquad\qquad CRITICAL REGION};
% FILL
\path[name path=xaxis]
(0,0) -- (\xmax,0);
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=0:\q}];
% LABELS
\node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$f(t|H_0)$};
\node[above right,black!20!red ] at ({1.05*(\S+\Ss)},{gauss(\S+\Ss,\S,\Ss)}) {$f(t|H_1)$};
\node[left, black!20!red, scale=1.3] at ({0.88*\q},{gauss(1.0*\q,\B,\Bs)}) {\strut$\beta$};
\node[right,black!20!blue,scale=1.3] at ({1.12*\q},{gauss(1.0*\q,\B,\Bs)}) {\strut$\alpha$};
\node[below=2pt] at ($(0,0)!0.4!(\q,0)$) {ACCEPT};
\node[below=2pt] at ($(\q,0)!0.6!(\xmax,0)$) {REJECT};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: p-value
\begin{tikzpicture}[inner frame sep=0]
\message{p-value^^J}
\def\q{5};
\def\B{3};
\def\S{8};
\def\Bs{1.0};
\def\Ss{1.5};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$t$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=250pt
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {gauss(x,\B,\Bs)};
%\addplot[name path=S,thick,black!10!red ] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q,{-0.03*gauss(\B,\B,\Bs)}) (\q, {4.0*gauss(\q,\B,\Bs)})}
node[below=-2pt,pos=0] {$t_\text{obs}$};
% FILL
\path[name path=xaxis]
(0,0) -- (\xmax,0);
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
% LABELS
\node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$f(t|H_0)$};
%\node[above right,black!20!red ] at ({1.05*(\S+\Ss)},{gauss(\S+\Ss,\S,\Ss)}) {$f(t|H_1)$};
\node[left,black!20!blue,scale=1.3] at ({0.98*\q},{0.5*gauss(\q,\B,\Bs)}) {$p$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: p-value, PLR
\begin{tikzpicture}
\message{p-value, PLR^^J}
\def\q{4.8}
\def\Bs{2.6}
\def\xmax{1.8*\q}
\def\ymin{{-0.16/\Bs}}
\def\ymax{{1.1/\Bs}}
\begin{axis}[every axis plot post/.append style={
mark=none,domain=0:{1.08*\xmax},samples=\N,smooth},
xmin={-0.05*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax=\ymax,
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q_0$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=240pt,
domain=0:(\xmax)
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {exp(-x/\Bs)/\Bs};
\addplot[black,dashed,thick]
coordinates {(\q,{-0.08*exp(-\q/\Bs)/\Bs}) (\q, {2.5*exp(-\q/\Bs)/\Bs})}
node[below=-2pt,pos=0] {$q_0^\text{obs}$};
% FILL
\path[name path=xaxis]
(0,0) -- (1.08*\xmax,0);
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:1.08*\xmax}];
% LABELS
\node[above right=2pt,black!20!blue] at ( 0.2*\Bs,{exp(-0.2)/\Bs}) {$f(q_0|\mu=0)$};
\node[left,black!20!blue,scale=1.3] at ({0.98*\q},{0.6*exp(-\q/\Bs)/\Bs}) {$p_0$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: p-value, PLR f(q|mu)
\begin{tikzpicture}
\message{p-value, PLR f(q|mu)^^J}
\def\q{4.8}
\def\S{1}
\def\Ss{\S*0.35}
\def\xmax{20}
\def\ymin{-0.05}
\def\ymax{0.5}
\begin{axis}[every axis plot post/.append style={
mark=none,domain=0:{1.08*\xmax},samples=\N,smooth},
xmin={-0.05*\xmax}, xmax=\xmax,
ymin=\ymin, ymax=\ymax,
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q_0$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=240pt,
domain=0:(\xmax)
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {fq0(x)};
\addplot[name path=B,thick,black!10!red] {fq(x,\S,\Ss)};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: expected sensitivity / p-value
\begin{tikzpicture}
\message{Expected sensitivty & p-value^^J}
\def\q{4.8}
\def\Bs{2.6}
\def\S{\q}
\def\Ss{1.4}
\def\xmax{1.8*\q}
\def\ymin{{-0.16/\Bs}}
\def\ymax{{1.1/\Bs}}
\begin{axis}[every axis plot post/.append style={
mark=none,domain=0:{1.08*\xmax},samples=\N,smooth},
xmin={-0.05*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax=\ymax,
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=240pt,
domain=0:(\xmax)
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {exp(-x/\Bs)/\Bs};
\addplot[name path=S,thick,black!10!red ] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q,{-0.08*exp(-\q/\Bs)/\Bs}) (\q, {1.08*gauss(\q,\S,\Ss)})}
node[above right=-2pt] {$\text{Med}[q_0|\mu]$}
node[below=-2pt,pos=0] {$q_0^\text{obs}$};
% FILL
\path[name path=xaxis]
(0,0) -- (1.08*\xmax,0);
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:1.08*\xmax}];
% LABELS
\node[above right=2pt,black!20!blue] at ( 0.2*\Bs,{exp(-0.2)/\Bs}) {$f(q_0|\mu=0)$};
\node[above right=2pt,black!20!red] at ({1.3*\q},{gauss(1.3*\q,\S,\Ss)}) {$f(q_0|\mu)$};
\node[left,black!20!blue,scale=1.3] at ({0.98*\q},{0.6*exp(-\q/\Bs)/\Bs}) {$p_0$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: bias & systematic error (variance)
\begin{tikzpicture}[inner frame sep=0]
\message{Bias & systematic error^^J}
\def\B{8};
\def\T{4};
\def\V{4};
\def\Bs{1.0};
\def\Ts{1.0};
\def\Vs{2.0};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\def\ymax{{1.1*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax=\ymax,
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$\hat{\theta}$,
ylabel={$f(\hat{\theta}|\theta)$},
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
every axis y label/.style={at={(current axis.above origin)},anchor=north east,align=center},
y=200pt
]
% PLOTS
\addplot[name path=B,thick,black!10!blue ] {gauss(x,\B,\Bs)};
\addplot[name path=S,thick,black!10!green] {gauss(x,\T,\Ts)};
\addplot[name path=S,thick,black!10!red ] {gauss(x,\V,\Vs)};
\addplot[black,dashed,thick]
coordinates {(\T,{-0.03*gauss(\T,\T,\Ts)}) (\T, {1.1*gauss(\T,\T,\Ts)})}
node[below=-2pt,pos=0] {$\theta$};
% % FILL
% \path[name path=xaxis]
% (0,0) -- (\xmax,0);
% \addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
%
% % LABELS
% \node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$f(t|H_0)$};
% %\node[above right,black!20!red ] at ({1.05*(\S+\Ss)},{gauss(\S+\Ss,\S,\Ss)}) {$f(t|H_1)$};
% \node[left,black!20!blue,scale=1.3] at ({0.98*\q},{0.5*gauss(\q,\B,\Bs)}) {$p$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: broadening from nuisance parameter
\begin{tikzpicture}
\message{Nuisance parameter^^J}
\def\B{3};
\def\S{8};
\def\Bs{1.0};
\def\Bbs{1.2};
\def\Ss{1.3};
\def\Sbs{1.5};
\def\xmax{\S+3.2*\Ss};
\def\ymin{{-0.15*gauss(\B,\B,\Bbs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper,
ticks=none,
xlabel=$t$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=250pt
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {gauss(x,\B,\Bs)};
\addplot[name path=B,dashed,thick,black!5!blue] {\Bbs/\Bs*gauss(x,\B,\Bbs)};
\addplot[name path=S,thick,black!10!red ] {gauss(x,\S,\Ss)};
\addplot[name path=S,dashed,thick,black!5!red ] {\Sbs/\Ss*gauss(x,\S,\Sbs)};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: p-value normal distributions
\begin{tikzpicture}[inner frame sep=0]
\message{Normal distrubution p-value^^J}
\def\q{5};
\def\B{3};
\def\S{8};
\def\Bs{1.0};
\def\Ss{1.5};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$x$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
y=250pt
]
% PLOTS
\addplot[name path=B,thick,black!10!blue] {gauss(x,\B,\Bs)};
%\addplot[name path=S,thick,black!10!red ] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q,{-0.03*gauss(\B,\B,\Bs)}) (\q, {4.0*gauss(\q,\B,\Bs)})}
node[below=-2pt,pos=0] {$x_\text{obs}$};
\addplot[black,dashed,thin]
coordinates {(\B,{-0.035*gauss(\B,\B,\Bs)}) (\B, {gauss(\B,\B,\Bs)})}
node[below=0pt,pos=0] {$\mu$};
\addplot[<->,black,thin]
coordinates {(\B,{gauss(\B-\Bs,\B,\Bs)}) (\B+\Bs, {gauss(\B+\Bs,\B,\Bs)})}
node[below,midway] {$1\sigma$};
\addplot[<->,black,thin]
coordinates {(\B,{2.6*gauss(\q,\B,\Bs)}) (\q,{2.6*gauss(\q,\B,\Bs)})}
node[below,midway] {$Z\sigma$};
% FILL
\path[name path=xaxis]
(0,0) -- (\xmax,0);
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
% LABELS
\node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$\mathcal{N}(\mu,\sigma)$};
\node[left,black!20!blue,scale=1.3] at ({0.98*\q},{0.52*gauss(\q,\B,\Bs)}) {$p$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: test statistics
\begin{tikzpicture}
\message{Test statistics^^J}
\def\q{6.3};
\def\B{8.3};
\def\S{4};
\def\Bs{1.0};
\def\Ss{1.5};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$t$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
width=0.7*\textwidth, height=0.5*\textwidth,
y=250pt
]
% PLOTS
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[red, name path=S,thick] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q, {0.95*gauss(\B,\B,\Bs)}) (\q, \ymin)}
node[below=3pt,anchor=south west] {$t_\text{cut}$};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=0:\q}];
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=\q:\xmax}];
% LABELS
\node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$f(t|H_0)$};
\node[above left=2pt,black!20!red] at (1.05*\S,{gauss(\S,\S,\Ss)}) {$f(t|H_1)$};
\node[below=2pt] at ($(0,0)!0.3!(\q,0)$) {REJECT};
\node[below=2pt] at ($(\q,0)!0.7!(\xmax,0)$) {ACCEPT};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: normal distributions, different mu
\begin{tikzpicture}
\message{Normal distributions, different mu^^J}
\def\q{5};
\def\B{3};
\def\S{7};
\def\Bs{1.0};
\def\Ss{1.0};
\def\xmax{\S+3.2*\Ss};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$x$,
x label/.style={at={(current axis.right of origin)},anchor=north west},
width=0.7*\textwidth, height=0.5*\textwidth,
clip=false, % prevent labels falling off
y=200pt
]
% PLOTS
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[red, name path=S,thick] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\B, {1.02*gauss(\B,\B,\Bs)}) (\B,{-0.05*gauss(\B,\B,\Bs)})}
node[below=-4pt] {\strut$\mu_0$};
\addplot[black,dashed,thick]
coordinates {(\S, {1.02*gauss(\S,\S,\Ss)}) (\S,{-0.05*gauss(\S,\S,\Ss)})}
node[below=-4pt] {\strut$\mu$};
\addplot[black,dashed,thick]
coordinates {(\q, {0.50*gauss(\B,\B,\Bs)}) (\q,{-0.05*gauss(\B,\B,\Bs)})}
node[below=-4pt] {\strut$x_\text{cut}$};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=0:\q}];
% LABELS
\node[above=2pt,black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$\mathcal{N}(\mu_0,\sigma)$};
\node[above=2pt,black!20!red] at (1.05*\S,{gauss(\S,\S,\Ss)}) {$\mathcal{N}(\mu,\sigma)$};
\node[right,black!20!blue,scale=1.3] at ({1.1*\q},{gauss(1.0*\q,\B,\Bs)}) {\strut$\alpha$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: power (cdf)
\begin{tikzpicture}
\message{Power (CDF)^^J}
\def\q{3};
\def\B{3};
\def\S{7};
\def\Bs{1.0};
\def\Ss{1.0};
\def\xmax{\S+2.0*\Ss};
\def\ymin{-0.15};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.05*cdf(\xmax,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$\mu$,
ylabel={power\\$1-\beta$},
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
every axis y label/.style={at={(current axis.above origin)},anchor=north east,align=center},
width=0.7*\textwidth, height=0.5*\textwidth,
clip=false, % prevent labels falling off
y=100pt
]
% PLOTS
%\addplot[name path=B,thick,black!10!blue] {cdf(x,\B,\Bs)};
\addplot[name path=B,thick,black!10!red] {1-cdf(\q,x,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\B,{1-cdf(\q,\B,\Ss)}) (\B,-0.05)}
node[below=-5pt] {\strut$\mu_0$};
\addplot[black,dashed,thick]
coordinates {(\B,{1-cdf(\q,\B,\Ss)}) (0,{1-cdf(\q,\B,\Ss)})}
node[left=0pt] {\strut$\alpha$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: low sensitivity 1
\begin{tikzpicture}
\message{Low sensitivity 1^^J}
\def\q{5.5};
\def\B{-1.0};
\def\S{-1.0};
\def\Bs{3.00};
\def\Ss{3.40};
\def\xmax{\S+3.2*\Ss};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
%axis x line=bottom, % no box around the plot, only x and y axis
%axis y line=left, % ...line*=... suppresses the arrow tips
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q_\mu$,
x label/.style={at={(current axis.right of origin)},anchor=north west},
width=0.7*\textwidth, height=0.5*\textwidth,
y=700pt
]
% plots
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[red, name path=S,thick] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q, {0.95*gauss(\B,\B,\Bs)}) (\q, \ymin)};
%node[below=2pt,anchor=south west] {$q_\text{obs}$};
\draw[->,thick]
(\q,{0.7*gauss(\B,\B,\Bs)}) -- ({\q+0.5*\Ss},{0.7*gauss(\B,\B,\Bs)})
node[below=8pt,align=center] {\qquad CRITICAL\\\qquad REGION};
% fill
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=\q:\xmax}];
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
% labels
\node[above right=2pt, black!20!blue]
at ( 0, {gauss(0,\B,\Bs)}) {$f(q_0|0)$};
\node[above right=2pt, black!20!red,align=center]
at ({\S+2.4*\Ss},{gauss(\S+2.4*\Ss,\S,\Ss)}) {$f(q_\mu|0)$};
%\node[above left, black!20!red ] at ({0.8*\q},{gauss(1.07*\q,\B,\Bs)}) {\strut$p_\text{s+b}$};
%\node[above right,black!20!blue] at ({1.1*\q},{gauss(1.07*\q,\B,\Bs)}) {\strut$p_\text{b}$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: low sensitivity 2
\begin{tikzpicture}
\message{Low sensitivity 2^^J}
\def\q{17.0};
\def\B{-1.0};
\def\S{18.0};
\def\Bs{10.0};
\def\Ss{8.0};
\def\xmax{\S+3.2*\Ss};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
%axis x line=bottom, % no box around the plot, only x and y axis
%axis y line=left, % ...line*=... suppresses the arrow tips
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q_\mu$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
width=0.7*\textwidth, height=0.5*\textwidth,
%y=700pt
]
% plots
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[red, name path=S,thick] {0.5*gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q, {gauss(\B,\B,\Bs)}) (\q, \ymin)};
\draw[->,thick]
(\q,{0.9*gauss(\B,\B,\Bs)}) -- ({\q+\Ss},{0.9*gauss(\B,\B,\Bs)})
node[below right=4pt,align=center] {CRITICAL\\REGION};
% fill
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=\q:\xmax}];
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=\q:\xmax}];
% labels
\node[above right=2pt, black!20!blue] at ( 0, {gauss(0,\B,\Bs)}) {$f(q_0|0)$};
\node[above right=2pt, black!20!red,align=center] at ({\S+1.1*\Ss},{0.5*gauss(\S+1.1*\Ss,\S,\Ss)}) {$f(q_\mu|0)$};
%\node[above left, black!20!red ] at ({0.8*\q},{gauss(1.07*\q,\B,\Bs)}) {\strut$p_\text{s+b}$};
%\node[above right,black!20!blue] at ({1.1*\q},{gauss(1.07*\q,\B,\Bs)}) {\strut$p_\text{b}$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: CLs, p-value
\begin{tikzpicture}
\message{CLs, p-value^^J}
\def\q{6.3};
\def\B{8.3};
\def\S{4};
\def\Bs{1.0};
\def\Ss{1.5};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
width=0.7*\textwidth, height=0.5*\textwidth,
y=250pt
]
% PLOTS
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[red, name path=S,thick] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q, {0.95*gauss(\B,\B,\Bs)}) (\q, \ymin)}
node[below=3pt,anchor=south west] {$q_\text{cut}$};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=0:\q}];
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=\q:\xmax}];
% LABELS
\node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$f(q|H_0)$};
\node[above left=2pt,black!20!red] at (1.05*\S,{gauss(\S,\S,\Ss)}) {$f(q|H_1)$};
\node[left, black!20!blue,scale=1.3] at ({0.9*\q},{gauss(1.04*\q,\B,\Bs)}) {\strut$p_\text{b}$};
\node[right,black!20!red, scale=1.3] at ({1.1*\q},{gauss(1.04*\q,\B,\Bs)}) {\strut$p_\text{s+b}$};
\end{axis}
\end{tikzpicture}
% GAUSSIANs: CLs, p-value large overlap
\begin{tikzpicture}
\message{CLs, p-value overlap^^J}
\def\q{6.8};
\def\B{6};
\def\S{4};
\def\Bs{1.0};
\def\Ss{1.5};
\def\xmax{\B+3.2*\Bs};
\def\ymin{{-0.15*gauss(\B,\B,\Bs)}};
\begin{axis}[every axis plot post/.append style={
mark=none,domain={-0.05*(\xmax)}:{1.08*\xmax},samples=\N,smooth},
xmin={-0.1*(\xmax)}, xmax=\xmax,
ymin=\ymin, ymax={1.1*gauss(\B,\B,\Bs)},
axis lines=middle,
axis line style=thick,
enlargelimits=upper, % extend the axes a bit to the right and top
ticks=none,
xlabel=$q$,
every axis x label/.style={at={(current axis.right of origin)},anchor=north west},
width=0.7*\textwidth, height=0.5*\textwidth,
y=250pt
]
% PLOTS
\addplot[blue, name path=B,thick] {gauss(x,\B,\Bs)};
\addplot[red, name path=S,thick] {gauss(x,\S,\Ss)};
\addplot[black,dashed,thick]
coordinates {(\q, {0.95*gauss(\B,\B,\Bs)}) (\q, \ymin)}
node[below=3pt,anchor=south west] {$q_\text{cut}$};
% FILL
\path[name path=xaxis]
(0,0) -- (\pgfkeysvalueof{/pgfplots/xmax},0); %\pgfkeysvalueof{/pgfplots/xmin}
\addplot[white!50!blue] fill between[of=xaxis and B, soft clip={domain=0:\q}];
\addplot[white!50!red] fill between[of=xaxis and S, soft clip={domain=\q:\xmax}];
% LABELS
\node[above=2pt, black!20!blue] at ( \B, {gauss(\B,\B,\Bs)}) {$f(q|H_0)$};
\node[above left=2pt,black!20!red] at (1.05*\S,{gauss(\S,\S,\Ss)}) {$f(q|H_1)$};
\node[below,black!20!blue,scale=1.3] at ({1.0*\B},{gauss(1.00*\q,\B,\Bs)}) {\strut$p_\text{b}$};
\node[right,black!20!red, scale=1.3] at ({1.05*\q},{gauss(0.95*\q,\S,\Ss)}) {\contour{white}{\strut$p_\text{s+b}$}};
\end{axis}
\end{tikzpicture}
\end{document}
Click to download: gaussians.tex • gaussians.pdf
Open in Overleaf: gaussians.tex



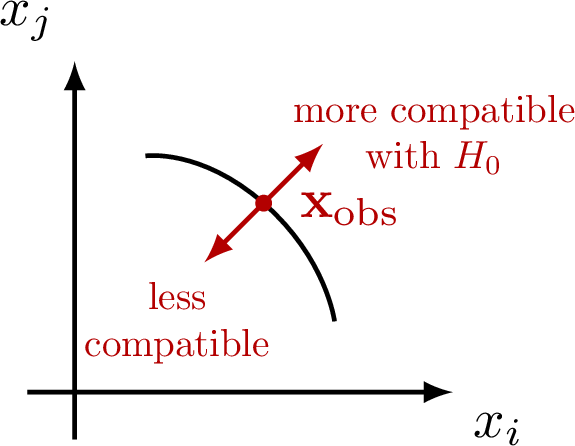
Hello,
Thank you very much for these very nice examples. I wanted to share the follow problem and fix for it: I downloaded the .tex file, and the figure for the 68-95-99 rule came out wrong. After some investigations I figured ou that in the lines that place the vertical dashed lines and that are of the form
\addplot[black,dashed,thick]
coordinates {({\B-3*\Bs},{20*gauss(\B-3*\Bs,\B,\Bs)}) ({\B-3*\Bs},{-\h})};
node[below=-3pt,scale=0.8] {\strut$\mu-3\sigma$};
the semi-colon on the second row should not be there… and it is not there in the editable code that one can play with directly on line. By removing the column, as follows, in all similar lines
\addplot[black,dashed,thick]
coordinates {({\B-3*\Bs},{20*gauss(\B-3*\Bs,\B,\Bs)}) ({\B-3*\Bs},{-\h})}
node[below=-3pt,scale=0.8] {\strut$\mu-3\sigma$};
the figure was again correct. The presence of the semi-colon does not produce a compilation error, but it seems that it messes up the coordinate system.